How to create a biological model on simBio
Dissolve the biological system
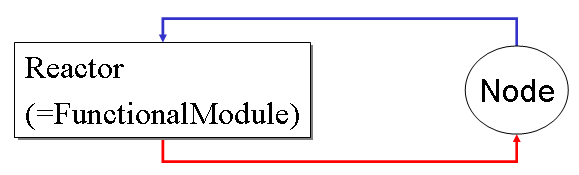
In simBio, the ordinary differential equations (ODE) are separated into a set of equations and variables according to the biological knowledge. A set of equations are defined as a Reactor, in other words, or functional module. Variables are defined as Nodes. A biological model is represented as a graph composed of Reactors and Nodes.
Write ODE in the Method "calculate(double)"
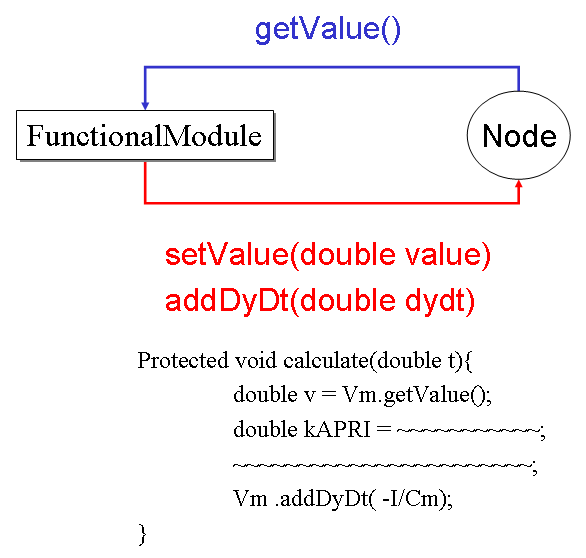
Create your Reactor which inherits core.Reactor and override Method "protected void calculate(double)". A set of equations are written in this Method. Node.getValue() is used to get the current value. Node.setValue(double) is used to set the current value. Node.addDyDt(double) is used to calculate ODE.
Define class field in Java source code
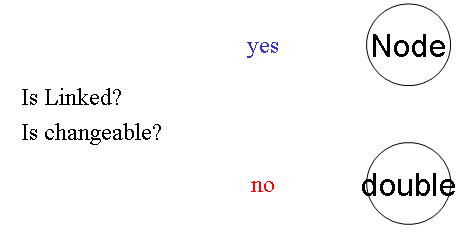
Nodes are separated into 2 groups in Java source code. One is "public Node" that is linked or calculated by ODE. Another is "public double" that is NOT linked from anywhere else, constant during integration. The name of the field must be the same as is in the XML file.
Write model description file in XML format
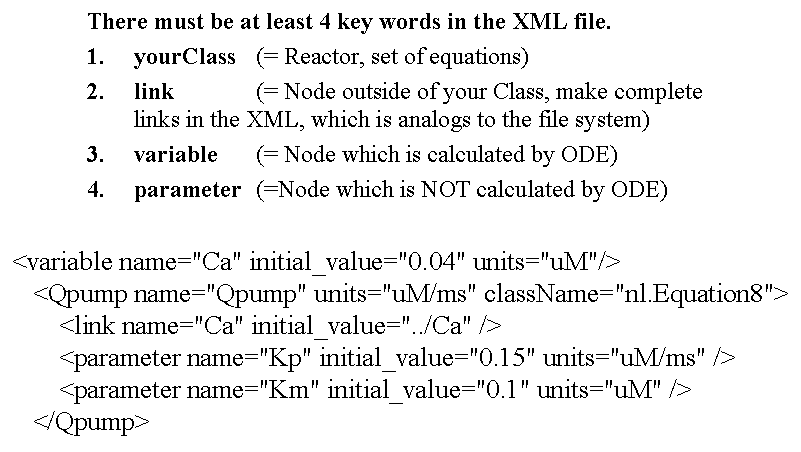

There must be at least 4 tags in the XML file.
- yourClass (= Reactor, set of equations)
- link (= Node outside of your Class, make complete links in the XML, which is analogous to the file system)
- variable (= Node which is calculated by ODE)
- parameter (=Node which is NOT calculated by ODE)
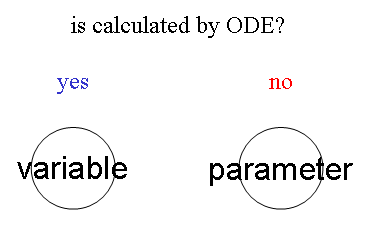
Nodes are separated into 2 groups. One is variable that is calculated by ODE. Another is parameter that is NOT calculated by ODE.

Link everything properly in the XML. The structure is analogous to the file system.
The core creates an instance tree and graph
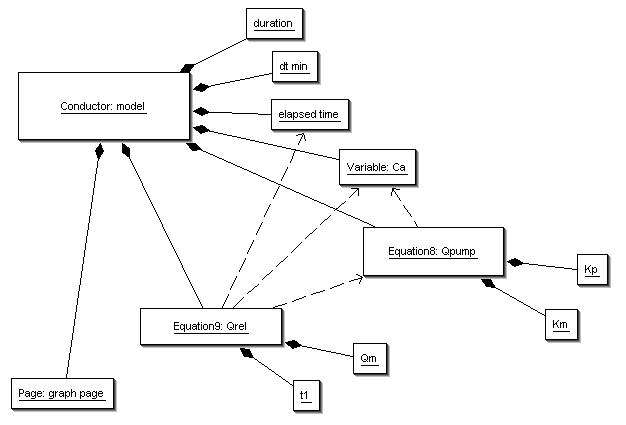
The core package creates an instance tree and automatically links everything as in the XML.
You may get the results
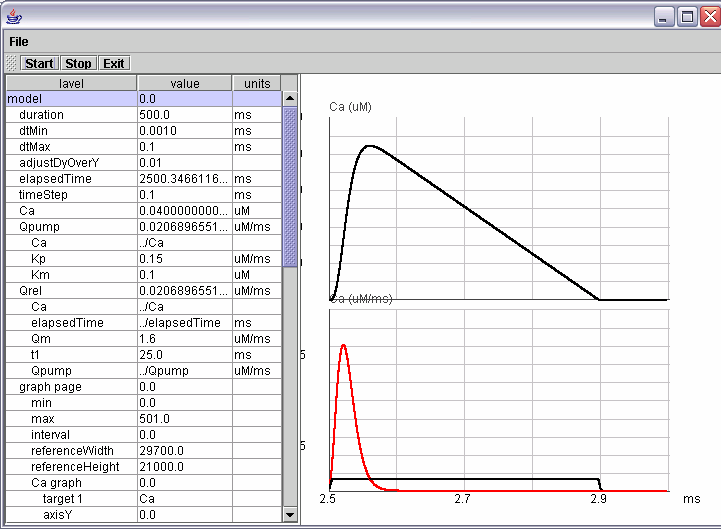
Every parameter can be edited and a 2D graph is available through the GUI.